Oligonucleotide therapeutics (e.g., antisense oligonucleotides (ASOs) and RNA interference (RNAi) agents) are revolutionizing treatment for hard-to-treat diseases, driven by their multi-target capability, high specificity, fast development cycles, and long-acting efficacy. Oligonucleotides have become an important strategy and have fueled a global surge in recent years [1,2].
For toxicokinetic (TK) and pharmacokinetic (PK) studies, reliable, high-throughput bioanalytical methods are critical for quantifying oligonucleotides and their metabolites in biological matrices (e.g., plasma, urine, and tissues). Current bioanalytical techniques include LC–MS/MS, fluorescence-based LC, immunoaffinity assays (e.g., ELISA), and qPCR. However, these methods exhibit distinct limitations: fluorescence-based LC lacks selectivity/ sensitivity for complex matrices; ELISA and qPCR, despite their high sensitivity, cannot distinguish full-length oligonucleotides from truncated metabolites, leading to overestimation of parent drug levels. In contrast, the LC-MS/MS method offers superior selectivity and simultaneous monitoring of target oligonucleotides and metabolites, making it the preferred choice for oligonucleotide bioanalysis [3]. For oligonucleotide bioanalysis, as in all methods, success depends on efficient sample preparation. This article reviews key sample preparation strategies for phosphorodiamidate morpholino oligomers (PMOs), ASOs, and small interfering RNA (siRNA), comparing principles, advantages, and applications (with case examples) to deliver an optimized LC-MS/MS bioanalytical workflow.
Key Challenges in LC–MS/MS Analysis of Oligonucleotides
Complex biological matrices, for example, blood, plasma, urine, and tissue, contain diverse components-salts, small organic molecules, lipids, phospholipids, proteins, and nonprotein macromolecules-all of which risk interfering with quantification. Compounding this, Oligonucleotides’ anionic charges lead to non-specific binding to matrix proteins and cell membranes. These factors create two critical challenges for LC–MS/MS analysis:
Matrix interference: Endogenous matrix components can compromise the accuracy of oligonucleotide quantification.
Low extraction recovery: Non-specific binding compromises conventional methods (e.g., protein precipitation), limiting extraction efficiency.
These hurdles call for a highly specific sample preparation method for reliable Oligonucleotide LC–MS/MS bioanalysis.
How to Choose Sample Preparation Methods for Oligonucleotide LC–MS/MS Analysis
Based on differences in the charged state under physiological conditions after phosphate-backbone modifications, those oligonucleotides can be categorized as:
Uncharged (nonionic phosphorodiamidate linkages, PMOs)
Negatively charged (negatively charged phosphate linkages, ASOs, and siRNA)
Uncharged oligonucleotides can typically be extracted effectively using protein precipitation (PPT). While negatively charged oligonucleotides, due to the strong protein binding, often necessitate liquid–liquid extraction (LLE), solid-phase extraction (SPE), hybridization extraction, or enzymatic digestion (Proteinase K digestion) combined with LLE or SPE [4, 5] to serve the low analyte recoveries. Several general sample preparation methods are described below.
Protein Precipitation (PPT) for Uncharged Oligonucleotides
Core Mechanism: A water-miscible organic solvent, for example, methanol or ethanol, is added to dehydrate proteins in the biological matrix, causing aggregation and precipitation. At the same time, analytes dissociate from proteins and partition into the organic phase. Sometimes, additives (e.g., guanidine hydrochloride, perfluoropropionic acid, buffers) are added before precipitation to accelerate the protein denaturation.
Advantages: Simple operation, controllable cost, and easy to implement.
Scope: Suitable for uncharged oligonucleotides(e.g., PMO).
Case Study: Using Viltolarsen as an example, methanol precipitation achieved a 70% recovery in rat plasma, with an LLOQ of 2 ng/mL (see Figure 1).
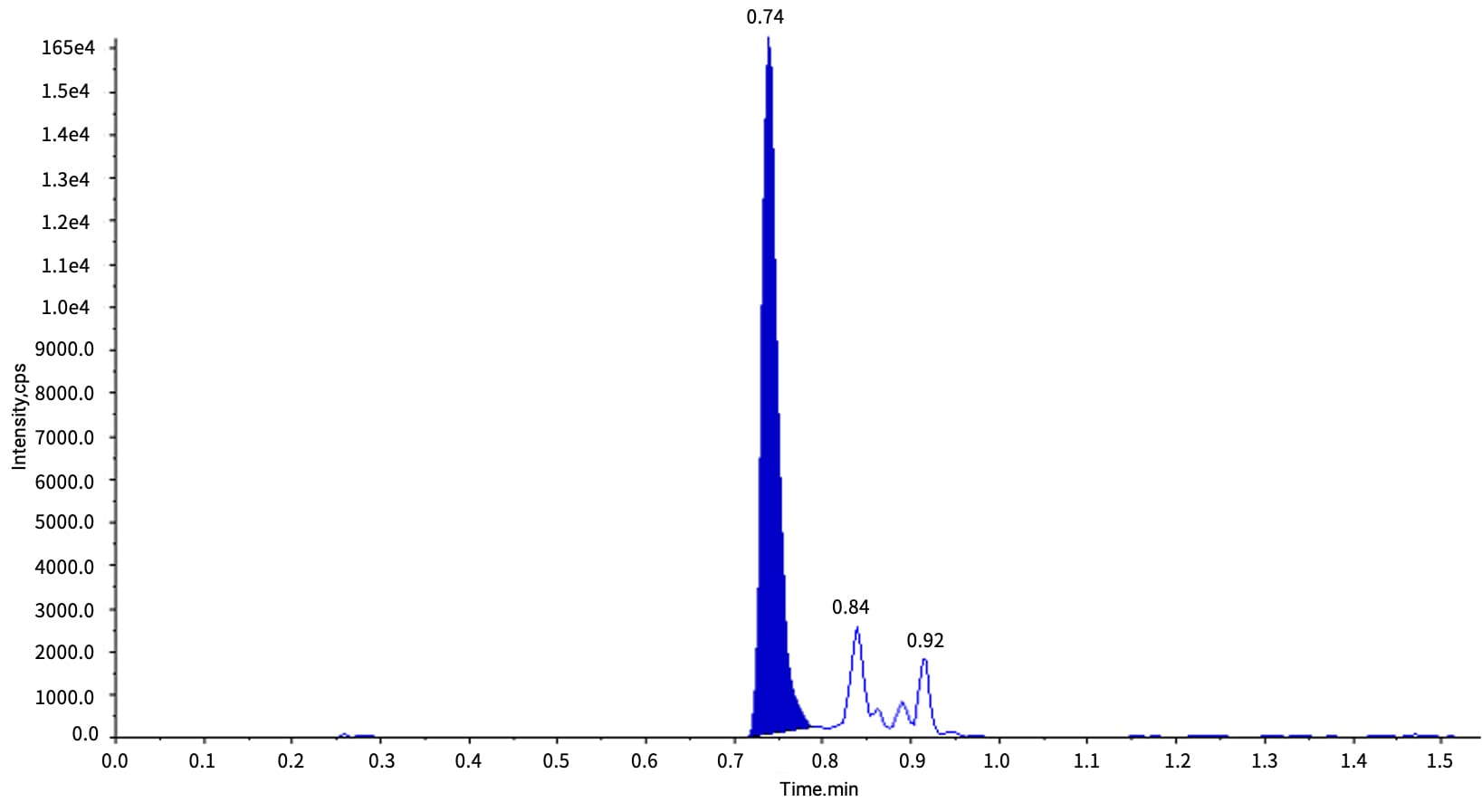
Figure 1. Typical chromatogram of 2 ng/mL Viltolarsen in rat plasma after PPT extraction
Liquid–Liquid Extraction (LLE) for Negatively Charged Oligonucleotides
Core Mechanism: A water-immiscible organic solvent mixture (phenol, chloroform, and isoamyl alcohol) is added to dissociate oligonucleotides from proteins and facilitate their transfer to the aqueous phase.
Phenol binds proteins to induce denaturation, releasing oligonucleotides to partition into the aqueous layer. Chloroform accelerates phase separation and extracts trace phenol from the aqueous phase, while isoamyl alcohol can minimize emulsification during extraction (see Figure 2).
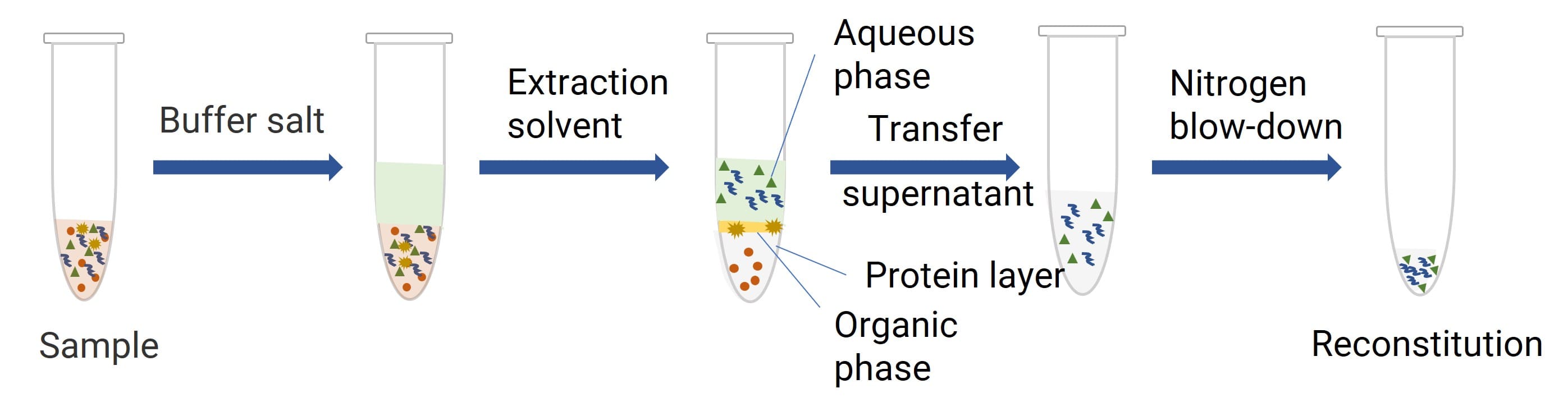
Figure 2. Schematic of LLE-Based Oligonucleotide Sample Preparation Workflow
Advantages: Standardized workflow with reproducible extraction recovery (>60%).
Scope: Suitable for hydrophilic, negatively charged analytes (e.g., siRNA, ASOs).
Limitations: Co-extraction of water-soluble endogenous components (e.g., salts, lipids, nucleic acids, peptides), and analogs elevates background, disrupts separation, and may induce matrix effects (ion suppression/enhancement), increased baseline noise, and variable recovery across matrices, especially for tissue homogenates and excreta (urine, bile, and feces). Plasma samples are basically unaffected.
Case Study:
LLextraction of Nusinersen (ASO) in plasma achieved an LLOQ of 2 ng/mL with >90% recovery (Figure 3).
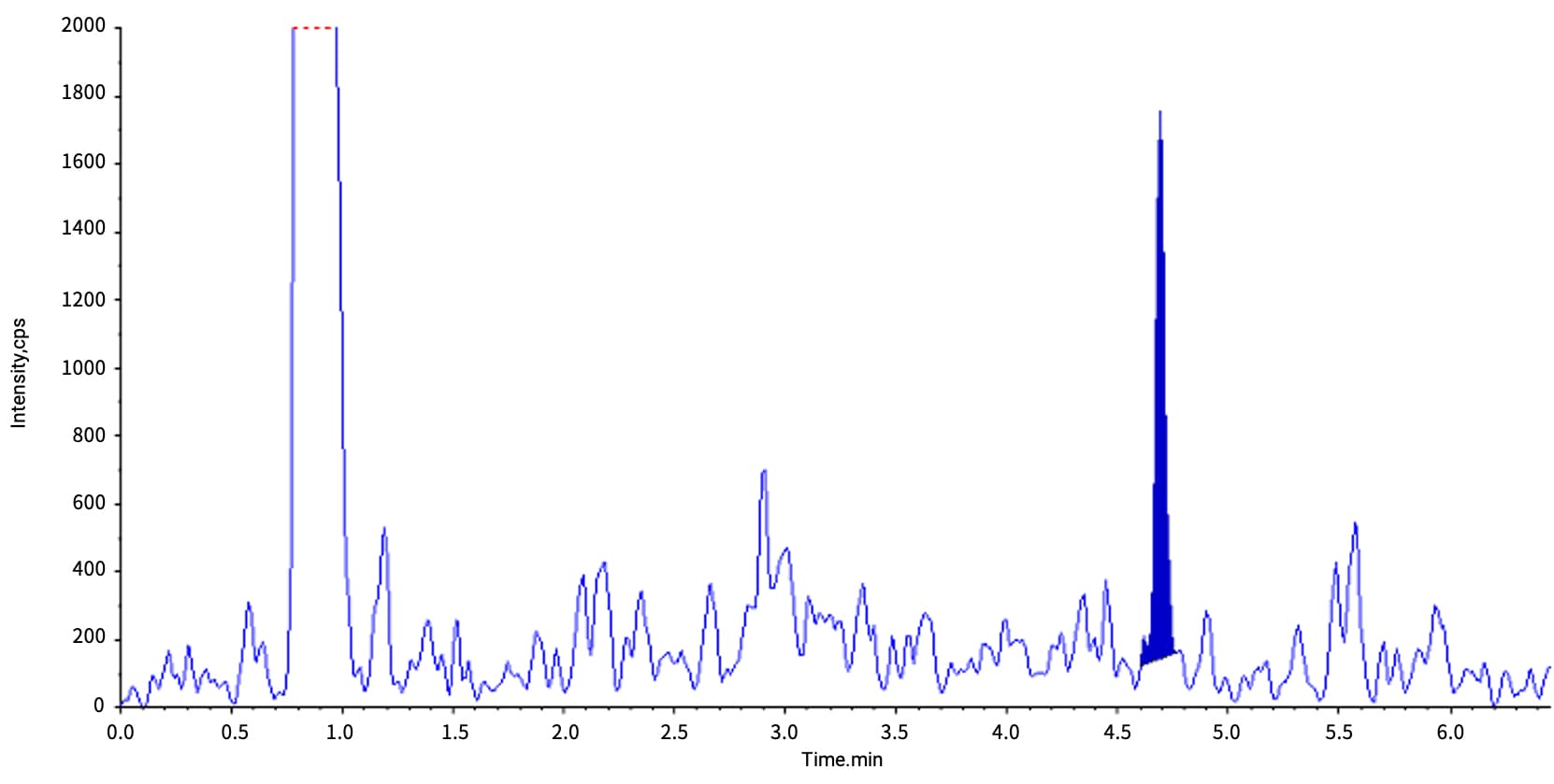
Figure 3. Typical chromatogram of Nusinersen at an LLOQ of 2 ng/mL after LLE extraction
During the measurement of Inclisiran (siRNA) in rat plasma and bile, the antisense (AS) strand exhibited a distinct signal across matrices. Matrix suppression was observed in bile, with the peak height of the 10 ng/mL rat plasma sample (retention time 3.82 min) being 7-fold higher than that of the bile sample (see Figures 4-1 and 4-2).
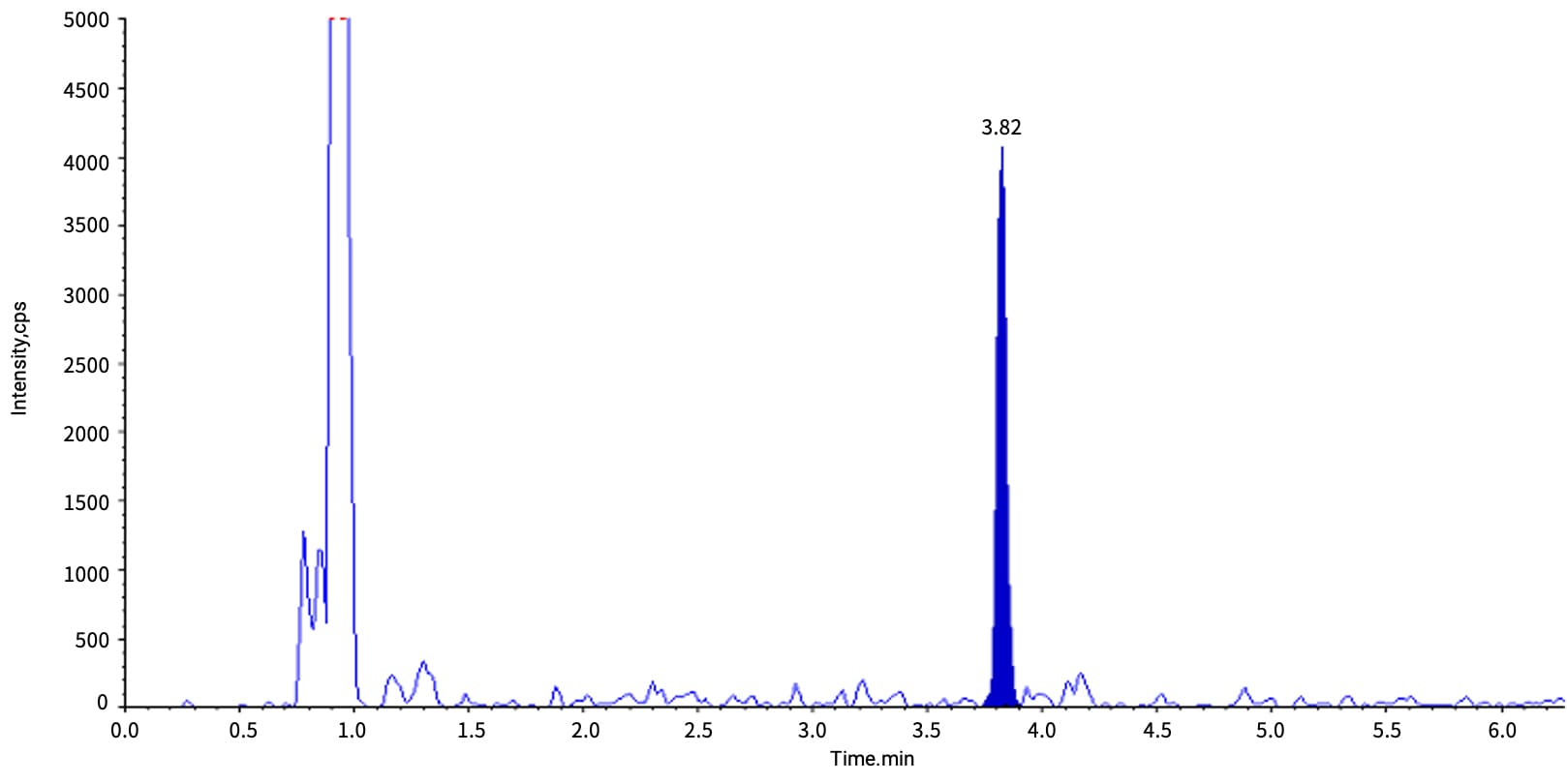
Figure 4-1. Typical chromatogram of Inclisiran-AS at 10 ng/mL in rat plasma matrix after LLE extraction

Figure 4-2. Typical chromatogram of Inclisiran-AS at 10 ng/mL in rat bile matrix after LLE extraction
Solid-Phase Extraction (SPE) for Reducing Matrix Effects and Enhancing Cross-Matrix Robustness
Core Mechanism: SPE isolates analytes by leveraging specific interactions with a solid sorbent: analytes are retained on the sorbent while interfering components are removed via washing. The target analyte is then eluted for downstream analysis (see Figure 6). For samples with strong analyte–protein binding, enzymatic digestion may be combined with SPE to improve extraction efficiency.
Key Retention Interactions
Hydrophilic–lipophilic balance (HLB) interaction: Ion-pairing reagents are added to form a neutral ion pair with the analyte, enabling binding to HLB sorbents (e.g., HLB cartridge, which balances hydrophilic and lipophilic properties).
Ion-exchange interaction: Buffer salts to dissociate the analyte into anionic forms, which then bind to anion-exchange sorbents such as WAX (weak anion exchange) cartridge.
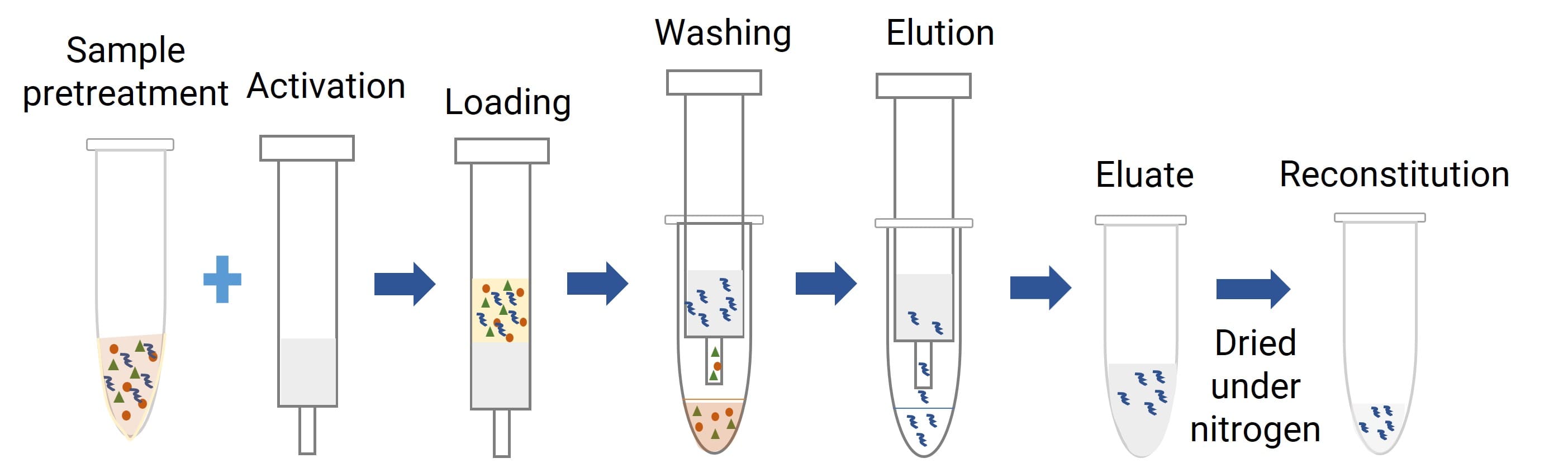
Figure 5. Schematic of SPE-Based Oligonucleotide Sample Preparation Workflow
Advantages: Enables cleanup and concentration, effectively reducing matrix interference. It also minimizes inter-matrix recovery variability and mitigates matrix effects.
Scope: Suitable for most oligonucleotide drugs and biological matrices that are stable under alkaline conditions, especially for oligonucleotides that show significant matrix-effect variability and suboptimal recovery across complex matrices (e.g., tissue homogenates, urine, bile, feces).
Case Study: For Inclisiran (siRNA), LLE-based isolation of the Inclisiran AS strand from rat bile yielded a low signal at 10 ng/mL (retention time of 3.83 min), accompanied by a prominent interference peak at 3.73 min. Switching to SPE improved sensitivity 10-fold and eliminated the interference peak (see Figures 6-1 vs. 6-2).
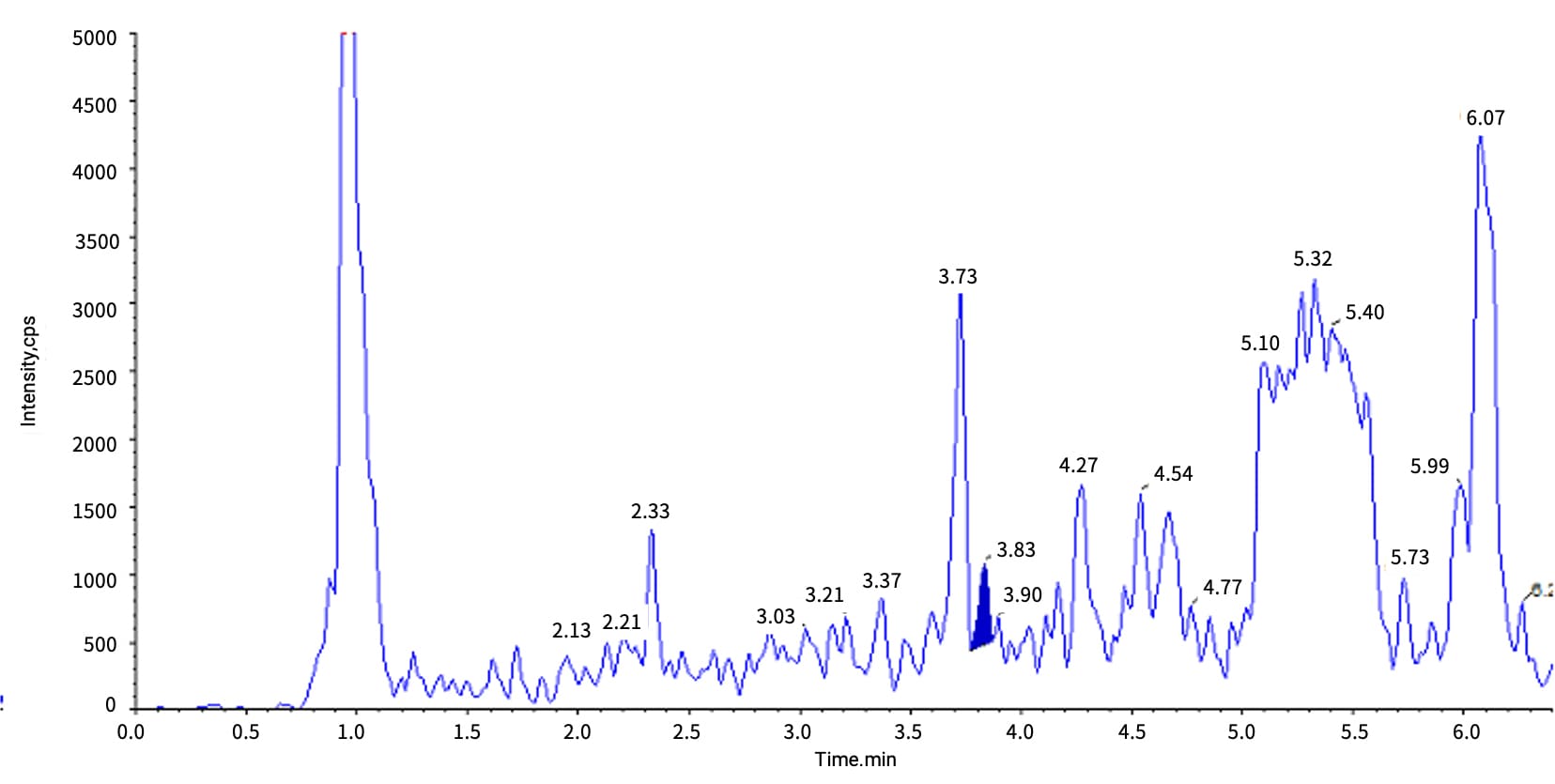
Figure 6-1. Typical chromatogram of Inclisiran-AS at 10 ng/mL in rat bile matrix after LLE extraction
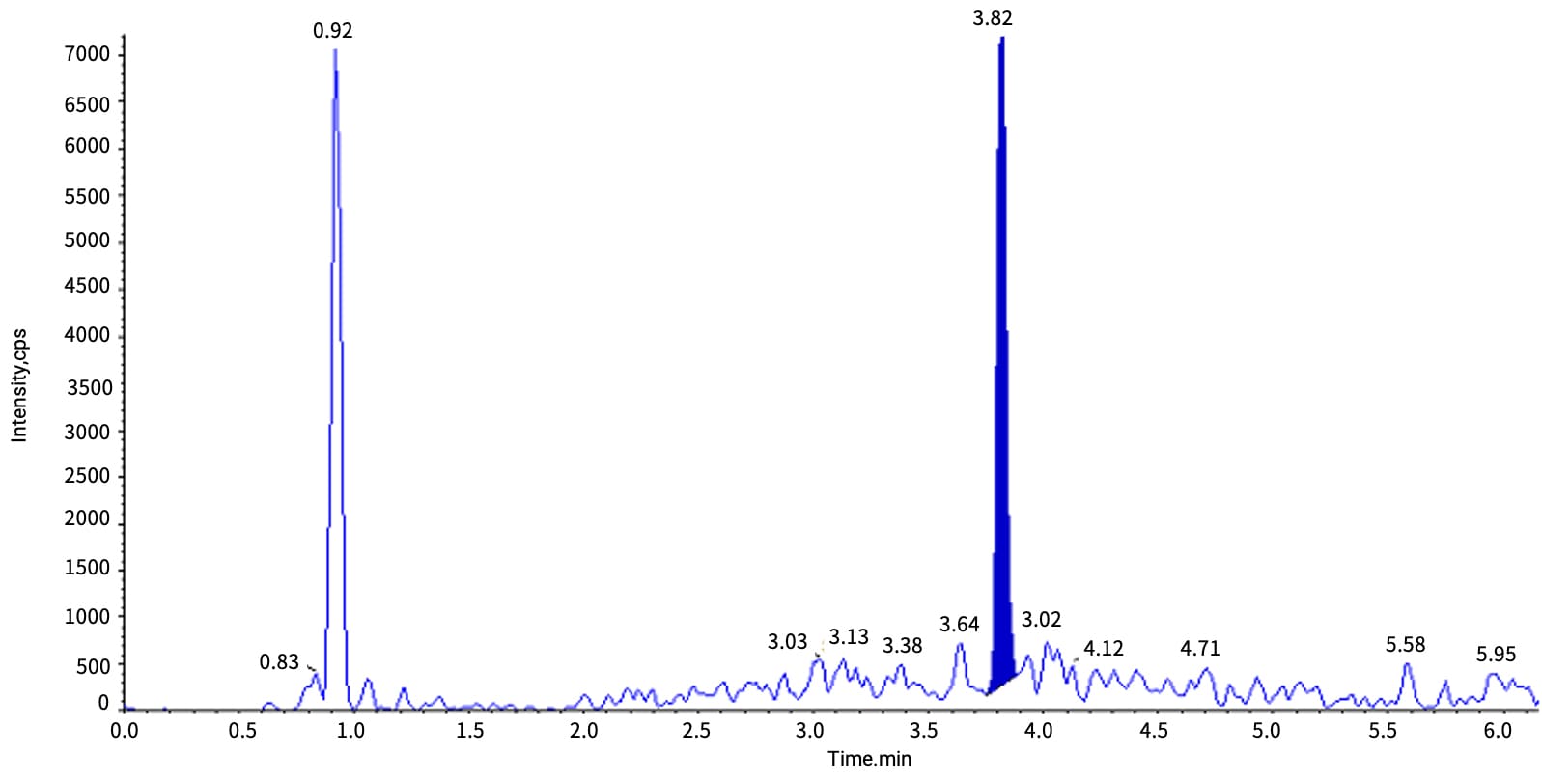
Figure 6-2. Typical chromatogram of Inclisiran-AS at 10 ng/mL in rat bile matrix after SPE extraction
Hybridization Extraction to Enhance Sensitivity and Specificity in Oligonucleotide LC–MS/MS Analysis
To improve sensitivity and minimize matrix background interference, hybridization-based LC–MS/MS methods have been developed in recent years. This target-specific approach requires a complementary biotinylated probe to enable highly specific and efficient extraction of the target oligonucleotide. Compared with traditional SPE or LLE, hybridization extraction significantly reduces matrix background, achieving sensitivity down to < 1 ng/mL.
As shown in Figure 7, the hybridization workflow: (1) Design a biotinylated probe based on the analyte’s base sequence. (2) Incubate the probe with streptavidin-coated magnetic beads to form probe-bead conjugates. (3) Add the conjugates to the biological matrix (pretreated by proteinase K) to capture the target oligonucleotide via complementary base pairing. (4) Discard the supernatant and wash the beads to remove unbound matrix components. (5) Release the analyte from the probe via high-temperature denaturation, yielding purified target analyte for downstream analysis [6].
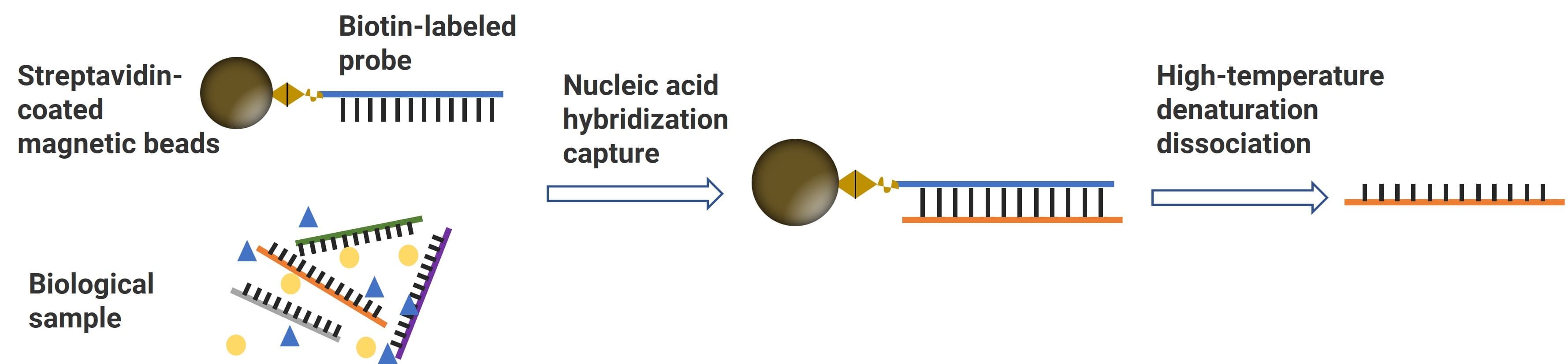
Figure 7. Schematic of nucleic-acid hybridization [5]
Advantages: High sensitivity, excellent selectivity.
Scope: Most oligonucleotide therapeutics.
Case study: For Inclisiran, typical generates two dominant product ions: m/z 78.9 (phosphate) and m/z 94.9 (phosphorothioate). In LC-MS/MS, the signal intensity of m/z 78.9 is generally over 5-fold higher than that of m/z 94.9. However, m/z 78.9, as a common fragment of endogenous nucleic acids, is frequently excluded as a product ion owing to high matrix background and interference. Following hybridization capture extraction from plasma, the matrix background was significantly reduced compared to SPE (see Figures 8-1 vs. 8-2).
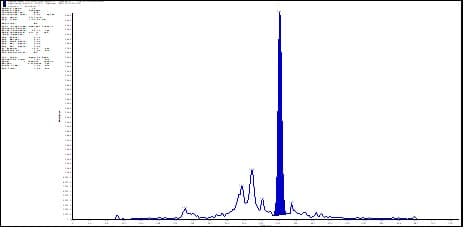
Figure 8-1. Typical chromatogram of Inclisiran-AS at 50 ng/mL in rat plasma matrix after SPE extraction
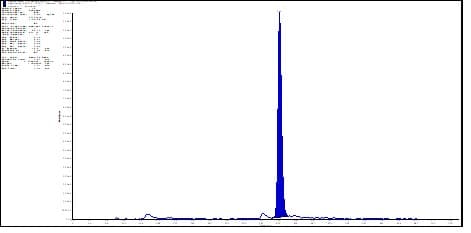
Figure 8-2. Typical chromatogram of Inclisiran-AS at 50 ng/mL in rat plasma matrix after hybridization extraction
Since analyte release from the probe depends on the melting temperature (Tm) of the probe-analyt hybrid complex, longer probes generally exhibit higher Tm values. However, high-temperature denaturation may also cause the bead-bound probes to dissociate from the beads. During chromatographic separation, the hybrid complexes may fail to dissociate at low column temperatures, while elevated column temperatures can reduce column longevity. Shorter probes exhibit lower Tm values but may compromise hybridization efficiency. Therefore, the optimal probe length requires experimental validation.
WuXi AppTec DMPK: Empowering Oligonucleotide Drug Discovery and Breaking Through Core Bottlenecks in Sample Preparation
In the development of oligonucleotide drugs, interference from complex biological matrices and the precise capture of ultra-low abundance targets remain key challenges restricting R&D efficiency. With profound insights into the molecular structures, chemical modifications, and mechanisms of action of oligonucleotide drugs, combined with over a decade of method development experience, WuXi AppTec DMPK Bioanalysis team has built an industry-leading mass spectrometry (MS) sample pre-processing technology platform. This platform enables flexible, customized solutions tailored to different oligonucleotide characteristics (e.g., sequence length, modification type), significantly shortening method development cycles while improving detection robustness and data reliability, thereby providing customers with high-quality research support.
To meet the full-cycle R&D needs of nucleic acid drugs, we provide comprehensive preclinical research services, ranging from in vitro ADME profiling and pharmacokinetic (PK) analysis to mass balance studies, as well as radiolabeling synthesis using radioisotopes (such as 3H, 14C) and supporting PK tracing.
WuXi AppTec DMPK is committed to accelerating the translation of your novel drugs from the laboratory to the clinic with professional and efficient services, and providing solid support for global partners in overcoming key R&D challenges.
Authors: Peiyun An, Jinlian Lu, Lili Xing
Talk to a WuXi AppTec expert today to get the support you need to achieve your drug development goals.
Committed to accelerating drug discovery and development, we offer a full range of discovery screening, preclinical development, clinical drug metabolism, and pharmacokinetic (DMPK) platforms and services. With research facilities in the United States (New Jersey) and China (Shanghai, Suzhou, Nanjing, and Nantong), 1,000+ scientists, and over fifteen years of experience in Investigational New Drug (IND) application, our DMPK team at WuXi AppTec are serving 1,600+ global clients, and have successfully supported 1,700+ IND applications.
Reference
[1] Jun Wang, Lan Wang. Efficacy analysis and research progress of marketed nucleic acid drugs. China New Drugs Journal, 2019, 28(18).
[2] Zhongzhe Cheng, Hongliang Jiang. Research progress in bioanalytical methods for nucleic acid drugs. Acta Pharmaceutica Sinica, 2021, 56(9): 2335–2345.
[3] William D van Dongen, Wilfried MA Niessen. Bioanalytical LC–MS of therapeutic oligonucleotides. Bioanalysis, 2011, 3(5), 541–564.
[4] Hyunsook Kim, Sujeong Kim. Oligonucleotide therapeutics and their chemical modification strategies for clinical applications. Journal of Pharmaceutical Investigation (2024) 54:415–433.
[5] Maria K. Tsoumpra, et al. Peptide-conjugate antisense-based splice-correction for Duchenne muscular dystrophy and other neuromuscular diseases. EBioMedicine 45 (2019): 630–645.
[6] Pei Li, Yuqing Gong, et al. Hybridization liquid chromatography–tandem mass spectrometry: An alternative bioanalytical method for antisense oligonucleotide quantitation in plasma and tissue samples. Analytical Chemistry, 2020, 92: 10548–10559.
Related Services and Platforms




-

 DMPK BioanalysisLearn More
DMPK BioanalysisLearn More -

 Novel Drug Modalities DMPK Enabling PlatformsLearn More
Novel Drug Modalities DMPK Enabling PlatformsLearn More -

 Novel Drug Modalities BioanalysisLearn More
Novel Drug Modalities BioanalysisLearn More -

 Small Molecules BioanalysisLearn More
Small Molecules BioanalysisLearn More -

 Bioanalytical Instrument PlatformLearn More
Bioanalytical Instrument PlatformLearn More -

 PROTAC DMPK ServicesLearn More
PROTAC DMPK ServicesLearn More -

 ADC DMPK ServicesLearn More
ADC DMPK ServicesLearn More -

 Oligo DMPK ServicesLearn More
Oligo DMPK ServicesLearn More -

 PDC DMPK ServicesLearn More
PDC DMPK ServicesLearn More -

 Peptide DMPK ServicesLearn More
Peptide DMPK ServicesLearn More -

 mRNA DMPK ServicesLearn More
mRNA DMPK ServicesLearn More -

 Covalent Drugs DMPK ServicesLearn More
Covalent Drugs DMPK ServicesLearn More
Stay Connected
Keep up with the latest news and insights.